This notebook demonstrates training the Neural Overlapping Community Detection model on a Facebook ego network dataset, reproducing the original interactive notebook workflow with the modernized codebase.
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from nocd import NOCD
from nocd.data import load_dataset
from nocd.metrics import overlapping_nmi, symmetric_jaccard, evaluate_unsupervised
from nocd.utils import plot_sparse_clustered_adjacency/home/runner/work/nocd/nocd/.venv/lib/python3.14/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Load the dataset¶
A(adjacency matrix):scipy.sparse.csr_matrixof size[N, N]X(attribute matrix):scipy.sparse.csr_matrixof size[N, D]Z_gt(ground truth communities):np.ndarrayof size[N, K]
graph = load_dataset('../data/facebook_ego/fb_698.npz')
A, X, Z_gt = graph['A'], graph['X'], graph['Z']
N, K = Z_gt.shape
print(f'Nodes: {N}, Features: {X.shape[1]}, Communities: {K}, Edges: {A.nnz}')Nodes: 66, Features: 6, Communities: 13, Edges: 540
/home/runner/work/nocd/nocd/.venv/lib/python3.14/site-packages/numpy/lib/_format_impl.py:838: VisibleDeprecationWarning: dtype(): align should be passed as Python or NumPy boolean but got `align=0`. Did you mean to pass a tuple to create a subarray type? (Deprecated NumPy 2.4)
array = pickle.load(fp, **pickle_kwargs)
Train the model¶
Using the scikit-learn compatible NOCD estimator with GCN + batch norm.
model = NOCD(
num_communities=K,
model_type='gcn',
hidden_dims=(64,),
batch_norm=True,
dropout=0.5,
lr=1e-3,
weight_decay=1e-2,
max_epochs=200,
display_step=50,
balance_loss=True,
stochastic_loss=True,
batch_size=5000,
)
model.fit(A, X, y=Z_gt)Using device: cpu
Epoch 0, loss = 0.5837, nmi = 0.2020
Epoch 50, loss = 0.4017, nmi = 0.1837
Epoch 100, loss = 0.3174, nmi = 0.2819
Epoch 150, loss = 0.2853, nmi = 0.2890
Epoch 200, loss = 0.2724, nmi = 0.3228
Training complete. Best NMI: 0.3228
Loading...
Evaluate¶
Z_pred = model.predict(A, X)
Z_soft = model.predict_proba(A, X)
nmi = overlapping_nmi(Z_pred, Z_gt)
jac = symmetric_jaccard(Z_pred, Z_gt)
unsup = evaluate_unsupervised(Z_pred, A)
print(f'Overlapping NMI: {nmi:.4f}')
print(f'Symmetric Jaccard: {jac:.4f}')
for k, v in unsup.items():
print(f'{k}: {v:.4f}')Overlapping NMI: 0.3228
Symmetric Jaccard: 0.2705
coverage: 0.8074
density: 0.6418
conductance: 0.2029
clustering_coef: 0.4011
Visualize the adjacency matrix sorted by communities¶
Left: predicted communities. Right: ground truth.
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
z_pred = np.argmax(Z_soft, axis=1)
o_pred = np.argsort(z_pred)
plot_sparse_clustered_adjacency(A, K, z_pred, o_pred, ax=axes[0], markersize=0.3)
axes[0].set_title('Predicted Communities', fontsize=14)
z_gt = np.argmax(Z_gt, axis=1)
o_gt = np.argsort(z_gt)
plot_sparse_clustered_adjacency(A, K, z_gt, o_gt, ax=axes[1], markersize=0.3)
axes[1].set_title('Ground Truth Communities', fontsize=14)
plt.tight_layout()
plt.show()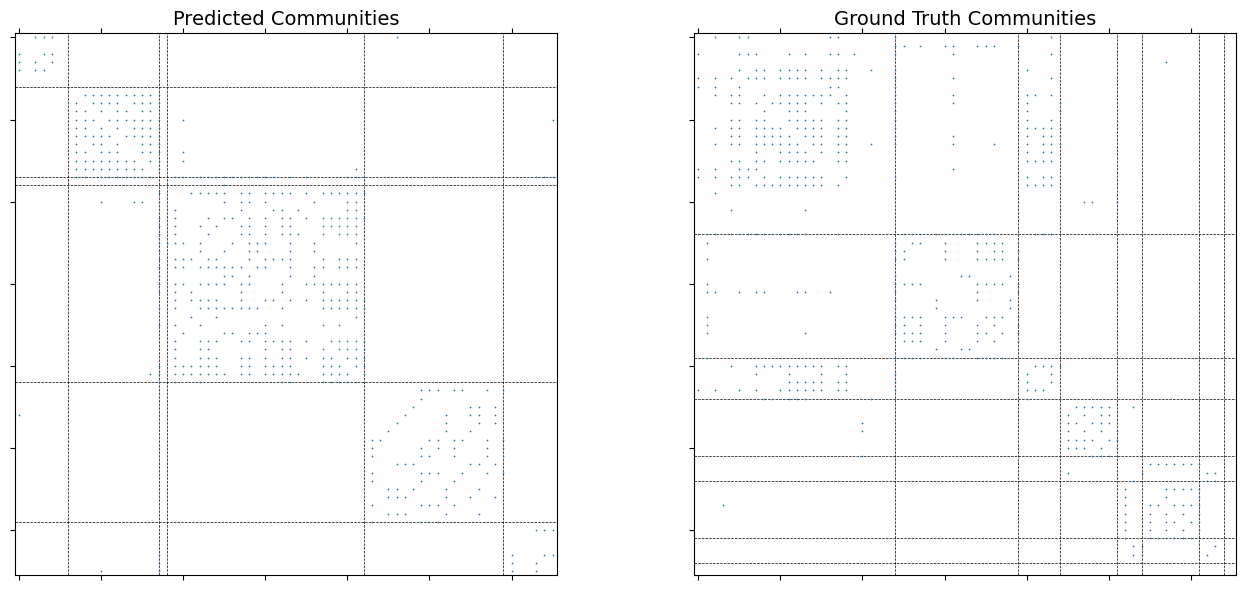
Save and reload¶
The model can be saved and loaded for later use.
model.save('/tmp/nocd_demo.pt')
loaded = NOCD.load('/tmp/nocd_demo.pt')
Z_loaded = loaded.predict_proba(A, X)
print('Predictions match:', np.allclose(Z_soft, Z_loaded, atol=1e-5))Predictions match: True